Scientist's Digital Notebook
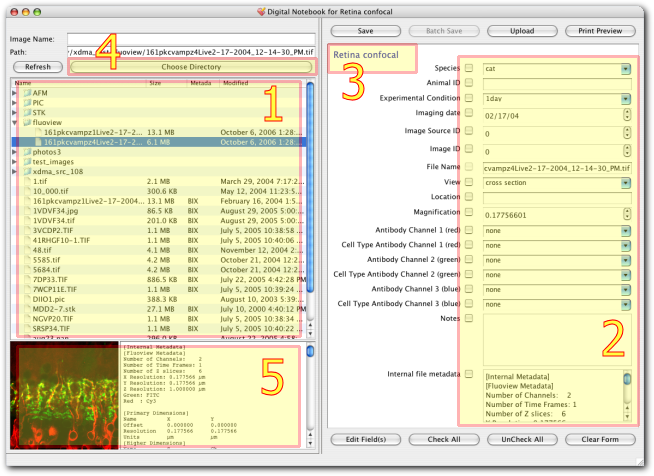
Digital Notebook is an application for biologists to manage image collections and related metadata both on their local machines and for integrating with the BISQUE database. It is designed to simplify the image annotation process and create required meta-data. Also it updates, prints and uploads existing information.
 Features
Features
- User-defined meta-data annotations
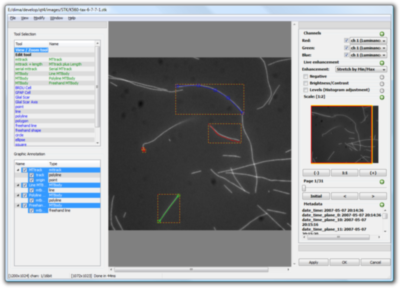
- Powerful graphical annotations with user-defined types
- Bio-formats meta-data and file name parsing
- Rapid upload for Bisque database
- In-place simple statistics for graphical annotations
Downloads
Documentation
- Quick Start: tutorial and configuration manuals
- Video Demos (see box on the right)
- Reference Paper: "Bisque: A Platform for Bioimage Analysis and Management"
- Changelog: version history
- Create Ticket: bug-reports/enhancements
Source Code
- Project Wiki: Trac timeline/tickets, Mercurial source code repository)
- Browse Source On-line
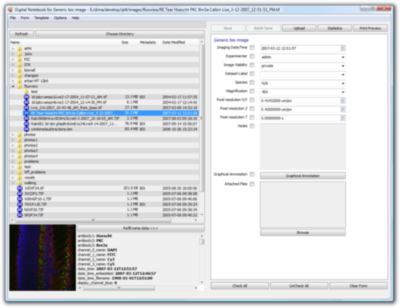
Digital Notebook is an application for biologists to manage image collections and related metadata both on their local machines and for integrating with the BISQUE database. The notebook eases the management and creation of metadata descriptions of large volumes of images by using domain knowledge to relieve the biologist of repetetive tasks. The metadata entry system is fully customizable to the specific need of an image set through an editable XML config file that stores information about what metadata fields are necessary for a certain class of images as well as default and common values for those fields. The motebook also has a built in file browser for easily navigating sets of images kept in separate folders. It can read JPEG, TIFF, and PNG formatted image files and stores the metadata in a companion XML file next to the image.
Among the many advanced features of the Digital Notebook are batch processing of images and their metadata, FTP uploading of files to the server, and automatic integration of the images and their metadata with the BISQUE database system. The Digital Notebook can also print out the metadata to a hard copy for keeping physical records in lab notebooks. The notebook has been through several iterations of refinement based on direct feedback from the biologists, helping to bridge the gap between microscope and database and thus enable higher data throughput. The notebook is built using the QT 4.1.0 open source library and binary versions are available for download for Windows and OSX in addition to the source code.